PROG0200 - Bacterial population
Consider a population of bacteria, each with a certain type of plasmid: $m$ with a plasmid type I, $n$ with a plasmid type II and $k$ with a plasmid type III. Every time two bacteria of a different plasmid type meet, their plasmid mutates to the subtraction of both types. For example, when a type I bacteria meets a type III bacteria, the plasmid mutates to a type II. Now we want to simulate the evolution of a given population throughout time.
Assignment
Develop a class Population of which every object represents a population of bacterias. Each with a plasmid of a certain type. All objects must have an attribute population: a list of numbers with value 1, 2 or 3, where every element of the list represents a bacteria from the population and indicates the corresponding value of which type the bacteria has (at a certain point in time). Furthermore, all object must also have an attribute encounters, in which the number of encounters between bacteria is kept track of. The class Population must also support the following methods:
- The initializing method __init__
must make sure the attribute population
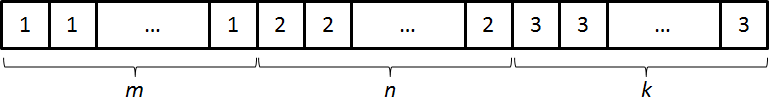
represents an initial population with $m$ bacteria with a plasmid type
I, $n$ with a plasmid type II and $k$ with a plasmid type III. The
values $m$, $n$ and $k$ are optional arguments of the method __init__
with standard value 0. After executing this method, the attribute encounters of the new object
must have value zero, and the attribute population
must have a value as is shown in the scheme below.
- The method plasmids must print a tuple ($m$, $n$, $k$) as a result, where the values $m$, $n$ and $k$ indicates the number of bacteria in the population with plasmids of respectively type I, type II and type III.
- The method __str__ must print a string of the format "type I: m, type II: n, type III: k (after x encounters)" as a result, where the values $m$, $n$, $k$ and $x$ must be filled out according to the current condition of the object.
- The method __repr__ must print a string of the format "Populatie(m, n, k)" as a result, where the values $m$, $n$ and $k$ must be filled out according to the current condition of the object.
- The method size must print the size of the population (total amount of bacteria) as a result.
- The method encounter
must simulate an encounter of two bacteria, as described in the
introduction. To this method, two parameters $0 \leq i,j < m+n+k$
must be given. If the bacteria $i$ and $j$ from the population have a
different plasmid type, the plasmid of both bacteria must mutate to the
third type. This is shown in the image below.
Make sure all methods are used optimally when implementing the class Population. The interactive Python session below illustrates the use of the class Population.
>>> population = Population(m=998, n=1, k=1) >>> print(population) type I: 998, type II: 1, type III: 1 (after 0 encounters) >>> print(repr(population)) Population(998, 1, 1) >>> print(population.plasmids()) (998, 1, 1) >>> for i in range(population.size()-1): ... population.encounter(i, i+1) >>> print(population) type I: 997, type II: 0, type III: 3 (after 999 encounters) >>> for i in range(population.size()-1): ... population.encounter(i, i+1) >>> print(population) type I: 997, type II: 3, type III: 0 (after 1998 encounters)
Now write another function
simulation(population[, number][, display)
to which as an obligatory argument population an object from the class Population must be given. This function must simulate a given number of encounters (optional argument number with standard value 1), where the two bacterias that meet are always chosen randomly from the given population. To follow up on the evolution of this population throughout these encounters, the function must print both the initial condition of the population and the condition of the population every so many encounters (given by the optional argument display with standard value 1). The printing of the condition of the population must happen conform the format of the method __str__ from the class Population. For example, the program code generates
population = Population(m=998, n=1, k=1) simulation(population, 10000, 250)
the following output (shortened)
type I: 998, type II: 1, type III: 1 (after 0 encounters) type I: 998, type II: 1, type III: 1 (after 250 encounters) type I: 995, type II: 4, type III: 1 (after 500 encounters) type I: 990, type II: 8, type III: 2 (after 750 encounters) ... type I: 337, type II: 357, type III: 306 (after 9500 encounters) type I: 328, type II: 348, type III: 324 (after 9750 encounters) type I: 337, type II: 324, type III: 339 (after 10000 encounters)
Remark: Because of the non-deterministic character of the output of the function simulation, the accurateness of your implementation of this function will not be assessed by Pythia. In the feedback given by Pythia, however, you can verify if the evolution of the population your code generates shows the same trend as the image below.
| Added by: | Peter Dawyndt |
| Date: | 2011-12-13 |
| Time limit: | 10s |
| Source limit: | 50000B |
| Memory limit: | 1536MB |
| Cluster: | Cube (Intel G860) |
| Languages: | PY_NBC |
| Resource: | None |


 RSS
RSS